After activation of CD4 + Th cells, they differentiate into functional Th1 and Th2 effector cells. Th1 cells secrete IL2 , IFNγ , TNFβ and other mediating cellular immune responses, delayed type hypersensitivity reactions and organ-specific autoimmune diseases in host anti-intracellular pathogens. It plays an important role in infection. Th2 cells produce IL4 , IL5 , IL6 , IL9 , IL10 , IL13 and other cytokine mediators, immune responses, allergic and infectious diseases in antagonizing extracellular pathogens ( such as bacteria, parasites) , B cell proliferation and differentiation, and asthma. Aspects play an important role. In recent years, extensive and in-depth research has been conducted on the mechanism by which CD4 + T cells differentiate into Th1 or Th2 cells. Some common understandings have been made: dual-signal stimulation activates CD4+ T cells, activation signals are transmitted intracellularly, and activation Multiple signal transduction pathways activate transcription factors, which in turn bind to the promoter region of the gene of interest, and promote the expression of protooncogenes, cytokine genes and their receptor genes. Cytokines can promote T cell clonal expansion through autocrine and paracrine effects , and then differentiate into functional Th1 and Th2 effector cells, and partially differentiate into memory cells. Cytokines are considered to be the most important factor in the differentiation of Th cells .
1 , TCR and co-stimulation signals
 Â
1.1TCR- CD3
The binding of TCR to the MHCII: antigen peptide complex molecule is the first signal required for T cell activation. After TCR2CD3 activation, CD3's Immune receptor tyrosine based activation motifs (ITAM) can transduce extracellular stimuli. In the early stage of T cell differentiation without IL4 and IL12, TCR activation signals influence the differentiation of Th1PTh2: MEKPERK activation, increased Ca2+ flux drives T cells to differentiate into Th1; PKC activation or partial blockade of calcineurin drives T The cells differentiate into Th2. In addition, weak TCR activation signal can activate Ca2+ flow signal to induce IL4 synthesis, and promote T cell differentiation to Th2; strong TCR activation signal can activate MAPK pathway to induce IFN2γ synthesis, and promote T cell differentiation to Th1. The length of TCR trigger time also affects the differentiation of Th cells; in the presence of IL12, a transient TCR triggers the initiation of Th1 differentiation, and a long-term TCR trigger initiates Th2 differentiation.
1.2 CD4
CD4 is a co-receptor of T cells, which binds to MHC class II molecules in the early stage of T cell activation and transduces signals through the protein tyrosine kinase LcK. LcK-deficient CD4+ T cells can exhibit Th2 differentiation disorder, Tec family kinase Rik And Itk (Lk downstream molecules) also play an important role in the Th2 differentiation process.
1.3 CD28 and its homologues
Binding of CD28 to B7.1 (CD80) or B7.2 (CD86) on activated APC provides a second signal for T cell activation. The production of IL4, IL5 and IL10 requires CD28 co-stimulation, and CD28 can increase the accumulation of the nuclear factor of activated T cells 2. In the absence of CD4 + T cells, only IFNγ is secreted. The APC experimental study of B7 knockout also confirmed that IL4 production and Th2 differentiation are highly dependent on B7, and experiments have found that CD28 can cause reorganization and sustained tyrosine phosphorylation of membrane lipid microdomains, but CD28 mediates Th2 differentiation. The molecular mechanism is still unclear. Recently, Skapenko et al. found that memory T cell CD28 can activate IL4 gene transcription and activate PI3 kinase, JNK-SAPK and p38 MAP pathway. CD28-B7 induces Th2 differentiation depending on IL4 expression, p38 and activation of ERK-MAP pathway. Therefore, CD8 may amplify the TCR activation signal that induces Th2 differentiation.
CTLA4 is a homolog of CD28, expressed in activated T cells, and binds to B7 with 10 times the affinity of CD28. In the absence of CTLA4, polyclonal T cell activation and proliferation disorder may occur. CTLA4 knockout T cells secrete high concentrations of IL4 and IL5, suggesting that CTLA4 down-regulates CD28 in the induction of Th2 differentiation.
The inducible costimulator (ICOS) is a recently discovered CD28 homologue that is also expressed in activated T cells. Unlike CD28 and CTLA4, ICOS does not bind to B7 but binds to B7RP1 (one of the homologs of B7). The IL4 secreted by T cells from ICOS-deficient mice was significantly reduced, while IFNγ was not abnormal. Therefore, ICOS can potentially regulate the differentiation and development of T cells.
2 , IL12
Many sources of IL4 can induce Th2 differentiation via STAT6, but it is unclear where the differentiated IL4 originally originated. The IL4 gene is located on chromosome 11, and there are IL5 and IL13 genes in this part. Several DNase I high-sensitivity sites associated with Th2 differentiation were found at the IL4 and IL13 loci, suggesting that chromatin recombination of these Th2-type cytokine gene loci is accompanied by Th2 differentiation. It has been confirmed that the hyperacetylation of histones on nucleosomes is associated with genes such as IL4, IL5, IL13, and this Th2-specific hyperacetylation is dependent on STAT6 and GATA3, accompanied by transcriptional expression of IL4 and IL13 genes. A conserved GATA3 response element (CGRE) sequence was found at 1. 6 kbp upstream of the IL13 gene locus, which binds to GATA3, histone acetyltransferase complex, and RNA polymerase II. It has the effect of enhancing the promoter of the above Th2 type cytokine gene. Through the study of the IL4 gene promoter, several transcription factors related to IL4 gene expression, such as NFAT2AP1, GATA3, cmaf and JunB, were also found. In addition, IL13 binds to the IL4 receptor alpha chain and regulates Th2 differentiation. IL26 up-regulates the expression of suppressor of cytokine signaling1 (SOCS1) and inhibits Th1 differentiation, while promoting the secretion of IL4 induced by activated CD4+ T cells. Differentiation.
3 , transcription factors
3.1 NF2AT
The NF2AT family has five members: NF2AT1 (NF2ATp), NF2AT2 (NF2ATc), NF2AT3, NF2AT4, and NF2AT5, all of which have highly conserved DNA and calcineurin binding sites. The first 4 are located in the cytoplasm and the NF2AT5 is located in the nucleus. Calcineurin phosphorylation mediates nuclear transport of NF2AT, and activation of protein kinases such as RasPRaf2MEK2ERK allows NF2AT to bind to AP21 to form a NFAT2AP1 complex that binds to DNA. Many gene promoters or enhancers expressed in activated T cells have NF2AT binding sites, such as IL2, IL4, IL25, IFNγ and the like. In mice knocked out of NF2AT2, expression of Th2 type cytokines such as IL4 was inhibited, presenting a defect in Th2 differentiation, and knocking out NF2AT1 or simultaneously knocking out NF2AT1 and NF2AT4 mice showed diametrically opposite results. This indicates that NF2AT1 and NF2AT2 bind to the NFAT2AP1 site of the IL4 promoter and function to regulate IL24 transcription in a different manner. Porter et al reported that ERK1, JNK3, and p38α are involved in the phosphorylation regulation of NF2AT and inhibit nuclear transport of NF2AT2.
3.2 GATA23
GATA23 belongs to the GATA family and is expressed in primary CD4+ T cells and Th0 cells that are differentiated into Th2 and is an important transcription factor for the regulation of Th2 differentiation. Transgenic mouse T cells overexpressing GATA23 can express Th2 type cytokine mRNAs such as IL4, IL5, IL6, IL10 and IL13, and inhibit GATA3 expression by antisense technology. The expression of these Th2 type cytokine mRNAs is also inhibited. However, there is no GATA23 binding site in the proximal promoter region of the IL4 gene, so it is thought that GATA23 may be a Th2-specific enhancer or exist as a local control region (eg, in the IL4, IL5, IL13 regions).
In STAT6 knockout T cells, GATA23 also induced Th2 type cytokine expression, leading to Th2 differentiation, and found a Th2-specific DNase I high-sensitivity site at the IL4 gene localization; meanwhile, GATA3 was able to bind to NF2AT The activity of the IL5 promoter. Recently, a structure molecule called FOG (Friend of GATA) has been discovered, which is expressed in the initial Th cells and acts to inhibit GATA23.
 Â
3.3 c2maf
C2maf is a basic region of the leucine zipper transcription factor, which is specifically expressed in Th2 cells and is the first cloned Th2-specific transcription factor. It has been thought that c2maf does not affect the expression of other Th2 type cytokines after activation, but induces IL24 expression to promote Th2 differentiation by selectively binding to the IL24 proximal promoter. Recently, researchers at Harvard Medical School found that c2maf also promotes Th2 differentiation via IL22Rα (CD25)-mediated but not IL4-dependent pathways. When c2maf is inactivated or knocked out, Th2 differentiation and secretion of corresponding cytokines proceed as usual, presumably related to the compensation of IL13.
3.4 T-bet
T-bet (also known as T-box 21) activates IFNγ expression and is a newly discovered Th1-specific transcription factor. T-bet can induce IFN-γ allele chromatin recombination and transcriptional activation of IFN-γ gene, and also induce IL12Rβ2 subunit expression, which promotes Th1 differentiation. When T2bet is introduced into differentiated CD4 Th2 cells, IFN2γ secretion is enhanced, Th2 type cytokines such as IL4 and IL5 are blocked, and cells are differentiated toward Th1. The study found that T2bet-induced Th1 differentiation is not dependent on IL12-PSTAT4 or IL18, but is inhibited by IL4-PSTAT6. However, the relationship between STAT and T2bet is currently unclear.
3.5 CPEBP and JunB
Similar to c2maf, members of the CPEBP family (such as CPEBPβ) also bind to the IL4 promoter and are potential inducers of endogenous IL4 expression. At the same time, CPEBP also inhibits the synthesis of IL22 and IFN-γ RNA, but does not affect other Th2 cytokines. JunB belongs to the c-jun transcription factor family and is selectively expressed in Th2 effector cells, which specifically binds to AP21 and activates IL-4 transcription. JunB-mediated IL-4 expression requires the cooperation of JNK kinase and c2maf and induces IL-2, IL-6 and IL-10 production in differentiated Th2 cells.
4. Signaling pathway in Th cell differentiation
4.1 JAK-STAT
JAK-STAT is a major signal transduction pathway mediated by cytokine receptors. IL-12 can mediate Th1 cell differentiation via STAT4, and IL-4 mediates Th2 cell differentiation via STAT6. However, the STAT-mediated gene of interest is still unclear, and no STAT binding site was found near the IL-4 promoter. Although STAT4 or STAT6 knockout experiments showed that IFN-γ or IL-4 were still secreted normally, the expression of GATA23, T-bet and cytokines was greatly affected by the lack of STAT signaling. It is speculated that STAT is closely related to transcription factors such as GATA23, c2maf and T2bet. Elucidating these intrinsic links may help to understand the molecular mechanism of Th cell differentiation. Bcl26 is considered to be a negative regulator of Th2 due to its ability to compete with STAT6 for the same binding site. It has recently been discovered that SOCS5 can negatively regulate the IL-4PSTAT6 pathway through interaction with IL-4R, thereby inhibiting Th2 differentiation.
4.2 MAPK
The mammalian MAPK signaling pathway includes four pathways: extracellular signal-regulated kinase (ERK), c-Jun N-terminal kinase (JNK) P stress-activated protein kinase (SAPK), P38MAPK, and ERK5PBMK1. Each pathway activates transcription factors through a conserved tertiary enzymatic cascade [MAPKKK→MAPKK(MKK)→MAPK], which regulates the expression of specific genes. ERK is located downstream of the oncogene Ras, and different activation signals such as Ras activate ERK via the Raf2MEK1 pathway, commonly referred to as the Ras2ERK pathway, including ERK1 and ERK2. It has been previously mentioned that the TCR activation signal activates MEKPERK and promotes Th1 differentiation. H2Ras transgenic mice have found that Ras2ERK pathway can up-regulate JAK1 kinase activity and promote STAT6 tyrosine phosphorylation after activation by non-TCR activation signal, suggesting a cross between Ras-ERK and JAK-STAT pathway. It can be seen that the exact relationship between the Ras2ERK pathway and Th cell differentiation remains to be studied.
There are four subtypes of p38 protein: p38-α, p38-β, p38-γ, and p38-δ. MKK3 and MKK6 are the major protein kinases that can be activated under stress or inflammatory stimuli. The direct substrate for p38 activation is MAPK-activated protein kinase (MAPKAPK) 22P3, which contains a CRE or a response element that is sensitive to interaction with the CREBPATF and AP21 families. Transgenic animals such as p382α and MKK3P6 were found to inhibit the activation of the p38MAPK pathway, and the level of IFN2γ was significantly decreased. Due to the presence of the c-junPATF2 binding site and a series of other ATF binding sites at the proximal and distal functional activation elements of the IFN2[gamma] promoter, and the proximal IFN2[gamma] element carrying the c-junPATF2 binding site, exhibits ThI specificity. Therefore, the p38MAPK pathway may induce Th1 cell differentiation by activating IFN2γ transcription.
JNK is also a stress-activated protein kinase, including JNK1P2P3. The activated phosphorylation cascade is MEKK1, 2 → MKK4PSEK1 → JNKPSAPK (MKK7 also activates JNK). JNKPSAPK can phosphorylate c2jun, ATF22 and increase its transcriptional activity, and promote the expression of c-fos, c-Jun and ATF22 regulatory genes. The JNK2 knockout study found that the level of IFN2γ in mice was decreased, and the Th1 immune response was defective, but the Th2 immune response was not affected. Addition of IFN-γ to JNK2-deficient T cells restored the above defects and produced a normal Th1 immune response. JNK1 knockout mice showed a strong Th2 immune response, and even under Th1 conditions, a large number of Th2 type cytokines such as IL-4, IL-5 and IL10 were produced, and the level of NF-AT2 in the nuclear of JNK1 knockout mice was found to increase. It is suggested that JNK1 can negatively regulate the level of NF2AT2 in the nucleus, thus affecting the production of Th2 type cytokines.
4.3 NF-κBPI-κB
Under normal conditions, NF2κB and its inhibitor I-κB are present in the cytosol. With the activation of the upstream kinase (NF-κB-inducible kinase or MEKK1), I-κB kinase is activated and I-κB protein phosphorylation results in their isolation and degradation from NF-κB, while free NF2κB is transferred into the nucleus. Transduction activates the gene of interest. The role of NF-κB factor in Th cell differentiation is just beginning. Studies have shown that inhibition of NF2κB activity blocks GATA23 expression and Th2 type cytokine production.
5, the conclusion
The process of cell differentiation is essentially the result of the sequential expression and selective expression of cellular genes. Therefore, the basic rule of cell differentiation regulation is the regulation of gene expression. For Th cell differentiation, the final expression is the activation and expression of genes involved in the control of cellular responses, and the cell signal transduction pathway plays an important mediating role here. In recent years, the research on the mechanism of Th cell differentiation has progressed rapidly. It has been recognized that factors such as TCR activation signal, co-receptor and cytokine environment mediate the differentiation direction of Th1PTh2 cells, and gradually deep into transcription factors, signal transduction, and genes. Regulatory level. We believe that the final clarification of the complexity of Th cell differentiation, the accuracy of regulation, and the cross-linking of signaling pathways and related factors will definitely bring about major breakthroughs in drug therapy for immune diseases.
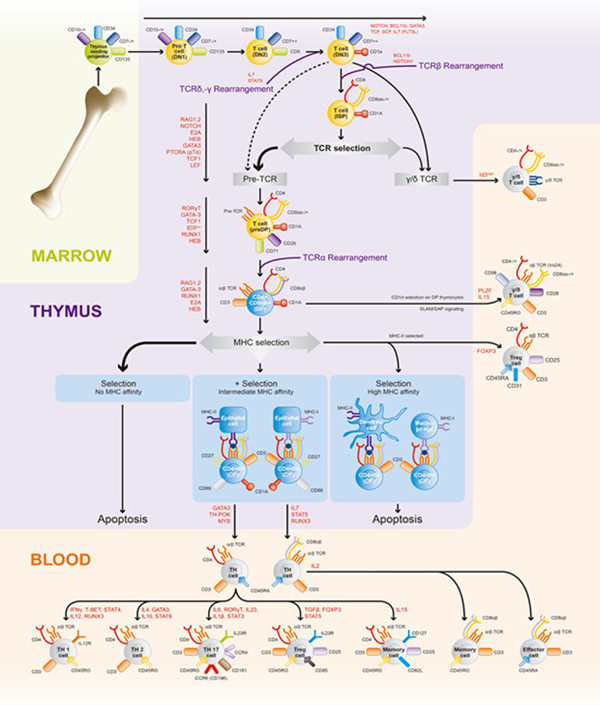
Related pathway indicators:
T-Cell Activation:
Regulators of T-Cell Activation: CD2, CD276, CD47, DPP4, CD3D, CD3E, CD3G, CD4, CD7, CD80, CD86, CD8A, CD8B, FOXP3, ICOSLG, IRF4, LAG3, LCK, MAP3K7 (TAK1), MICB, NCK1, TNFSF14, VAV1.
T-Cell Proliferation: CD28, CD3E, ICOSLG, IL1B, IL10, IL12B, IL18, NCK1, RIPK2, TNFSF14.
T-Cell Differentiation: ADA, APC, BCL2, BLM, CD1D, CD2, CD27 (TNFRSF7), CD4, CD80, CD86, EGR1, IL12B, IL15, IL2, IRF4, NOS2 (iNOS), PTPRC, SOCS1.
T-Cell Polarization: CCL3, CCR1, CCR2, CCR3, CCR4, CCR5, CD274, CD28, CD4, CD40LG (TNFSF5), CSF2 (GM-CSF), CXCR3, CXCR4, IFNG, IL12A, IL12RB1, IL12RB2, IL18R1, IL2 , IL4, IL4R, IL5, TGFB1.
Regulators of Th1 and Th2 Development: CD2, CD40 (TNFRSF5), CD5, CD7, CSF2 (GM-CSF), IFNG, IL10, IL12A, IL13, IL3, IL4, IL5, TLR2, TLR4, TLR9.
Th1 & Th2 Differentiation: CD28, CD40 (TNFRSF5), CD40LG (TNFSF5), CD86, IFNG, IL12A, IL12B, IL12RB1, IL12RB2, IL18, IL18R1, IL2, IL2RA, IL4, IL4R, IL6.
B-Cell Activation:
Antigen Dependent B-cell Activation: CD28, CD4, CD40 (TNFRSF5), CD40LG (TNFSF5), CD80, FAS (TNFRSF6), FASLG (TNFSF6), IL10, IL2, IL4.
Other Genes involved in B-Cell Activation: ADA, CXCR5, ICOSLG, IL6, IL7, MS4A1, TGFB1.
B-Cell Proliferation: BCL2, CD27 (TNFRSF7), CD40 (TNFRSF5), CD81, IL10, IL7, PTPRC.
B-Cell Differentiation: ADA, AICDA, BLNK, CD27 (TNFRSF7), IL10, IL11, IL4, RAG1.
Other Immune Cell Activation:
Macrophage Activation: IL13, IL4, TLR1, TLR4, TLR6.
Neutrophil Activation: IL8.
Natural Killer Cell Activation: CD2, IL12A, IL12B, IL2.
Leukocyte Activation: CX3CL1.
Pathway research tools:
Cell lineage identification series | PCR Array of Cell Lineage Identification | HU-CLI-084 |
Insulin resistance series | PCR Array of Insulin Resistance | HU-INR-084 |
T cell and B cell activation series | PCR Array of T-Cell & B-Cell Activation | HU-TCA-084 |
Terminal differentiation indicator series | PCR Array of Terminal Differentiation Markers | HU-TDM-084 |
PCR Array :
Q-PCR array is an important tool for basic biological research and clinical disease research in signaling pathways. This functional classification chip uses mature SYBR Green fluorescence quantitative PCR technology to focus on the detection of gene expression levels in a signaling pathway. Compared with traditional high-density expression profiling chips, Q-PCR array has the advantages of high pertinence, high sensitivity, accuracy and reliability. The genes on the chip include genes that have a definite relationship with the subject, or at least the genes whose relationship with the subject is to be verified. If the subject is a gene of a certain biological pathway, it is more efficient and convenient to use a functional classification gene chip for the gene than to use a high-density table to match the chip.
BioTNT QPCR chip features:
• High sensitivity and low sample usage;
• Wide linear range for simultaneous detection of genes with large differences in expression levels;
• Optimized detection primers for each gene to ensure high amplification rates and single product;
• High repeatability, the average difference in Ct values ​​is only 0.25 cycles;
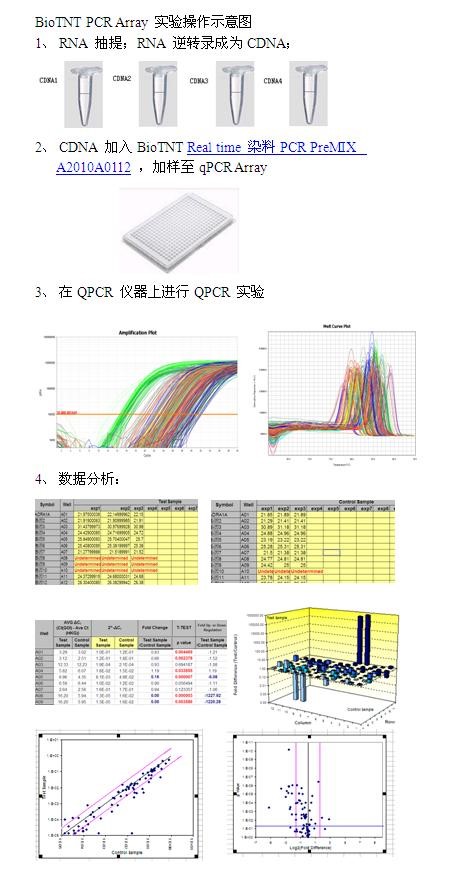
BioTNT QPCR Array chip operation process:
Urine Analyzer,Urine Analyzer For Lab,Lab Hospital Clinical Urine Analyzer,Urine Chemistry Analyzer
Guangdong Widinlsa International Co.Ltd , https://www.widinlsas.com